|
10/11/2023 0 Comments 10x chromium gel bead sequence
Specifically, i f the clog occurred near the end of the run, most cells will have been partitioned into GEMs. 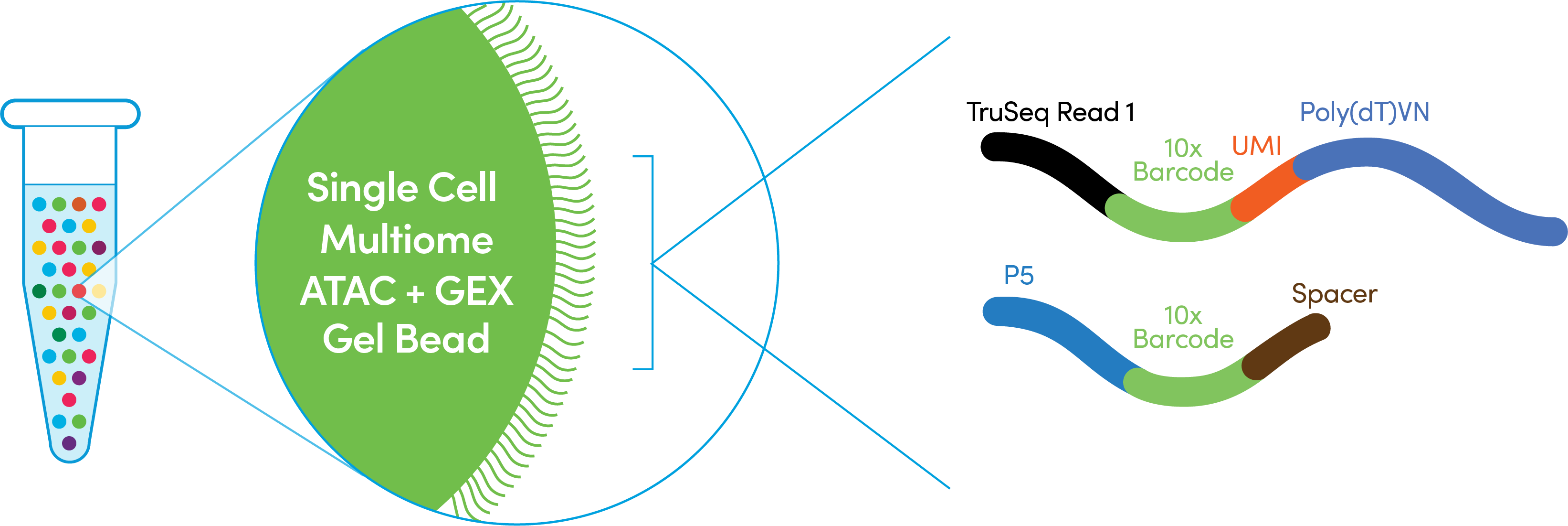 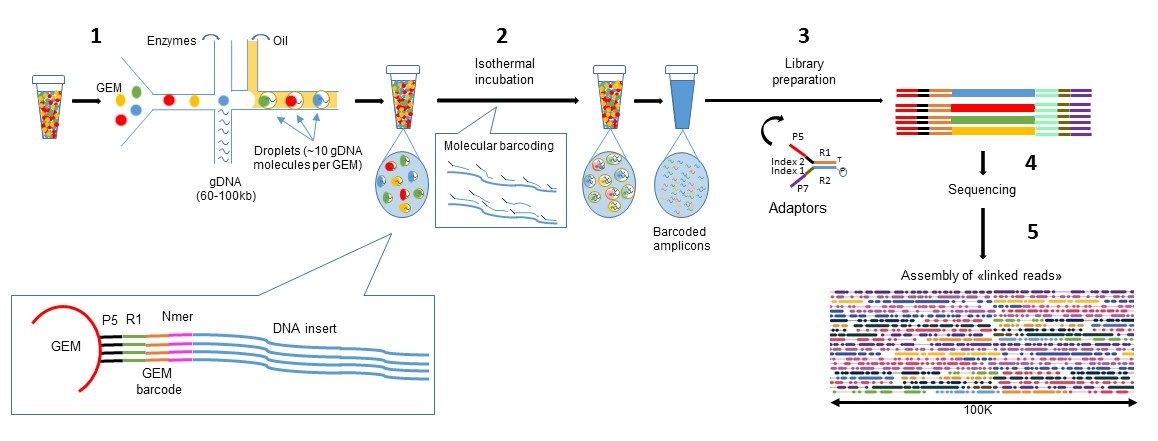
However, if it is a precious sample in which additional cell material cannot be obtained and cells would be lost otherwise, it may be possible to continue with the protocol and generate cDNA, albeit at lower levels, and with the knowledge that the number of recovered cells will be lower. We strongly recommend rerunning the sample if a clog occurs during GEM generation. Other materials other than pipette tip lids have not been tested, may introduce unwanted debris, and are not recommended. For HT chips (in which secondary holders are open while loading), a pipette tip lid may be placed over the chip itself and does not need to fully cover the opened portion with the gasket attached. To further minimize the rate of microfluidic clogs resulting from the introduction of dust / debris during chip loading, pipette tip lids may be used to cover the open chip between loading steps. To reduce the rate of microfluidic clogs, we recommend storing and loading chips in an area free of dust and debris. They will result in faster than usual depletion of Gel Beads and lower cDNA yield. Sample clogs are the result of suboptimal sample preparation of single-cell suspensions.A clog in the Gel Bead line causes Gel Beads to flow slower, leaving an excess of Gel Beads behind. Gel Bead clogs are caused by improper handling of Gel Beads.Best Practices to Minimize Chromium Next GEM Chip Clogs and Wetting FailuresĬlogs are generally caused by sub-optimal sample preparations, non-sterile work environments, clumping of Gel Beads and/or slow chip loading.ĭistinguishing between sample clogs and Gel Beads clogs:.If other samples are run in parallel, the clogged sample wells may have uneven volumes of remaining reagents compared to other sample wells.Īdditional details can be found in our Technical Note: This is evidenced by recovering less than 100 ul GEMs from the recovery well (see figure below) or an excess of Partitioning Oil in the recovered GEMs. Up to eight samples can be processed per batch within minutes.Question: Should I proceed if there is a sample clog?Īnswer: When running samples in the Chromium Controller, occasionally a sample clog may occur. The single-cell encapsulating process is significantly faster compared to inDrop or Drop-Seq. Drop-Seq) the 10X controller is capable of loading “all” droplets with micro-beads, enabling single-Poisson distribution loading and thus high capture efficiencies (in contrast to double-Poisson loading of other protocols). Each GEM is loaded with adapters containing one of 750,000 different barcodes for the single cell RNA-seq library preps. The flexible workflow encapsulates 500 to 80,000 cells or nuclei per sample together with gel beads to form gel beads in emulsions (GEMS). The technology allows for high-throughput single cell transcriptomics of a wide variety of cell types as well as single-nuclei expression profiling. The 10X Genomics Single Cell Chromium Platform is the single-cell expression profiling platform enabling the analysis of large cell numbers at the highest capture efficiency (of up to 65%).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
